THOIPApy
The Transmembrane HOmodimer Interface Prediction Algorithm (THOIPA) is a machine learning method for the analysis of protein-protein-interactions.
THOIPA predicts transmembrane homodimer interface residues from evolutionary sequence information.
THOIPA helps predict potential homotypic transmembrane interface residues, which can then be verified experimentally. THOIPA also aids in the energy-based modelling of transmembrane homodimers.
Important links:
How does thoipapy work?
- downloads protein homologues with BLAST
- extracts residue properties (e.g. residue conservation and polarity)
- trains a machine learning classifier
- validates the prediction performance
- creates heatmaps of residue properties and THOIPA prediction
Installation
pip install thoipapy
THOIPA has only been tested on Linux, due to reliance on external dependencies such as FreeContact, Phobius, CD-HIT and rate4site. For predictions only, a dockerised version is available that runs on Windows or MacOS. Please see the THOIPA webserver for the latest information.
Dependencies
We recommend the Anaconda python distribution, which contains all the required python modules (numpy, scipy, pandas,biopython and matplotlib). THOIPApy is currently tested for python 3.8.5. The requirements.txt contains a snapshot of compatible dependencies.
Development status
The code has been extensively updated and annotated for public release. However is released "as is" with some known issues, limitations and legacy code.
Usage as a standalone predictor
- first check if your needs are met by the THOIPA webserver or the latest version of dockerised software
- for local predictions on linux, first install phobius, NCBI_BLAST, biopython, freecontact, CD-HIT, and rate4site
- please see thoipapy/test/functional/test_standalone_prediction.py for the latest run syntax, typically
from thoipapy.thoipa import get_md5_checksum, run_THOIPA_prediction
from thoipapy.utils import make_sure_path_exists
protein_name = "ERBB3"
TMD_seq = "MALTVIAGLVVIFMMLGGTFL"
full_seq = "MVQNECRPCHENCTQGCKGPELQDCLGQTLVLIGKTHLTMALTVIAGLVVIFMMLGGTFLYWRGRRIQNKRAMRRYLERGESIEPLDPSEKANKVLA"
out_dir = "/path/to/your/desired/output/folder"
make_sure_path_exists(out_dir)
md5 = get_md5_checksum(TMD_seq, full_seq)
run_THOIPA_prediction(protein_name, md5, TMD_seq, full_seq, out_dir)Example Output
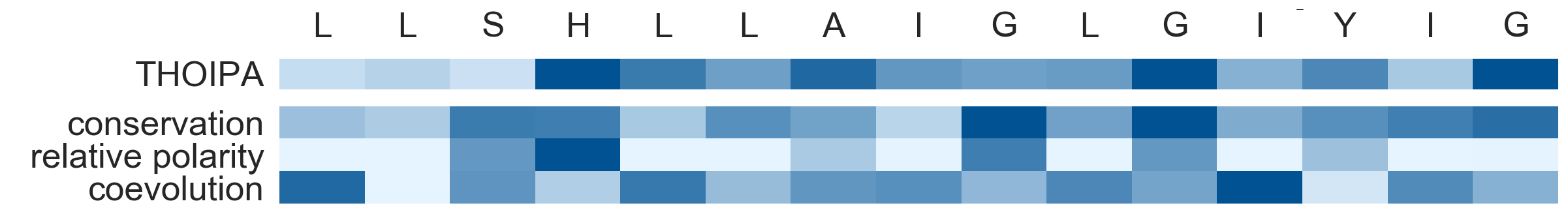
- the output includes a csv showing the THOIPA prediction for each residue, as well as a heatmap figure as a summary
- below is a heatmap showing the THOIPA prediction, and underlying conservation, relative polarity, and coevolution
Create your own machine learning predictor
- THOIPA can be retrained to any dataset of your choice
- the original set of training sequences and other resources are available via the Open Science Foundation
- the THOIPA feature extraction, feature selection, and training pipeline is fully automated
- contact us for an introduction to the THOIPA software pipeline and settings
python path/to/thoipapy/run.py -s home/user/thoipa/THOIPA_settings.xlsxLicense
THOIPApy is free software distributed under the permissive MIT License.
Contribute
- Contributors are welcome.
- For feedback or troubleshooting, please email us directly and initiate an issue in Github.
Contact
- Mark Teese, TNG Technology Consulting GmbH, formerly of the Langosch Lab at the Technical University of Munich
- Bo Zeng, Chinese Academy of Sciences, Beijing formerly of the Frishman Lab at the Technical University of Munich